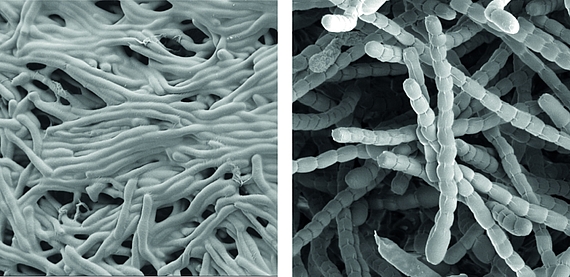
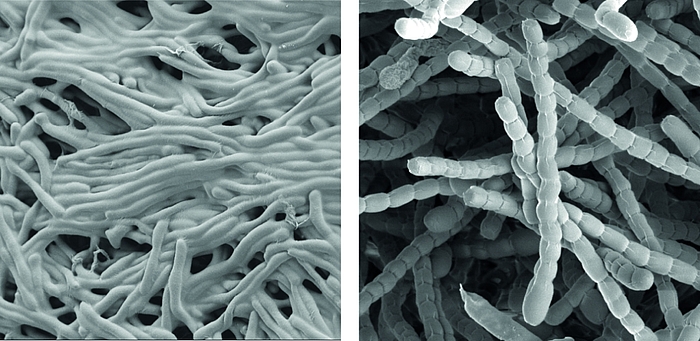
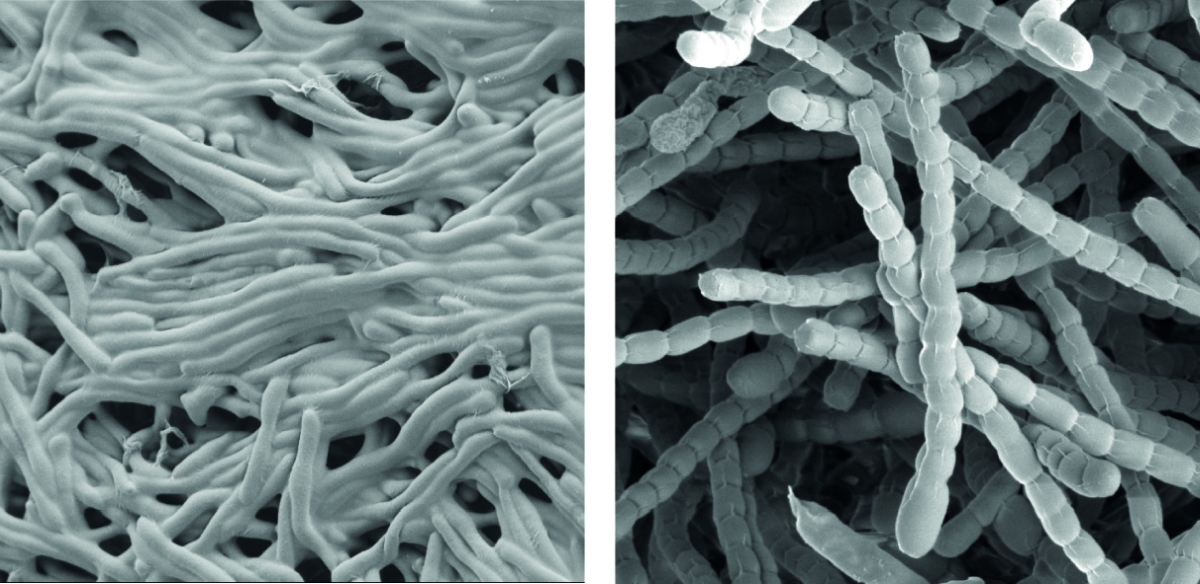
Streptomyceten sind wichtige ‘mikrobielle Fabriken’ von Wirkstoffen, sie produzieren die Mehrheit der heute in der Medizin verwendeten Antibiotika.
Als wahre Formwandler stellen sie auch ein faszinierendes Modell zur Erforschung der Zelldifferenzierung und bakterieller Multizellularität dar. Im Verlauf ihres Lebenszyklus durchlaufen sie eine komplexe Morphogenese von filamentösen, multizellulären, vegetativen Hyphen zu unizellulären, dormanten und hoch-resistenten Sporen.
Unser Team
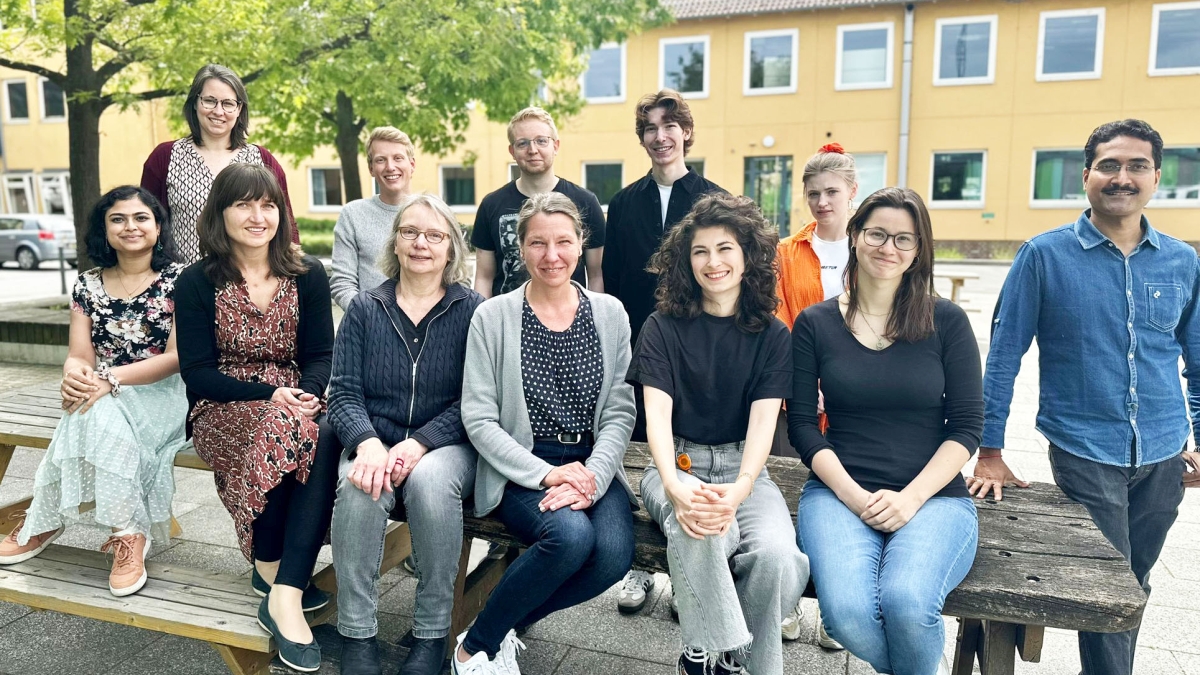



Möchtest auch Du Teil des Teams werden?
-
Informationen für PostdoktorandInnen, DoktorandInnen oder Masterstudierende
Wir freuen uns immer, neue PostdoktorandInnen, DoktorandInnen oder Masterstudierende in unserem internationalen Forschungsteam willkommen zu heißen. Hast Du Interesse an grundlegenden biologischen Fragestellungen in den Bereichen
• Signaltransduktion
• Stressadaptation
• Zelldifferenzierung
• Naturstoffe und Biopolymere
• Bakterielle Multizellularität
• Regulatorische Netzwerke und/oder
• Streptomyceten?
Hast Du Erfahrung oder Vorkenntnisse in Mikrobiologie, Genetik, Zellbiologie oder Biochemie? Dann sende bitte eine E-Mail an Natalia (tschowri@ifmb.uni-hannover.de) mit Deinem Lebenslauf und einem Motivationsschreiben. Fördermöglichkeiten ergeben sich oft kurzfristig, daher sind Initiativbewerbungen sehr willkommen!
Forschungsprojekte
Zyklisches di-GMP-abhängige Signaltransduktion und Regulation in bakterieller Zelldifferenzierung
Der sekundäre Botenstoff zyklisches di-GMP kommt in beinahe allen Bakterien vor und ist als ein zentraler Regulator zwischen motiler und sessiler Lebensweise in den meisten Bakterien anerkannt. In den nicht-motilen Streptomyceten reguliert zyklisches di-GMP die Transition zwischen der filamentösen Existenzform und den Sporen. Hohe c-di-GMP Mengen bewirken eine Blockade der Entwicklung, sodass die Zellen in ihrem vegetativen Stadium verharren. Niedrige c-di-GMP Mengen haben einen gegensätzlichen Effekt und führen zur verfrühten Sporulation. Unsere Studien führten zu der Entdeckung, dass BldD, der Masterregulator der Streptomyces Differenzierung, als ein c-di-GMP-Effektor fungiert. Bindung des tetrameren c-di-GMP an BldD führt zur Dimerisierung und somit Aktivierung von BldD als DNA-bindendes Protein und hat eine Repression von Sporulationsgenen zur Folge.
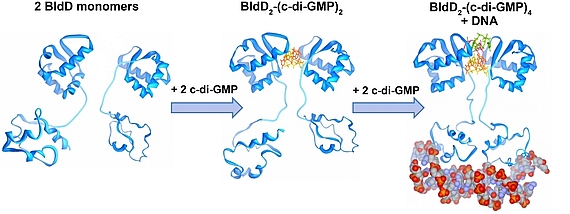
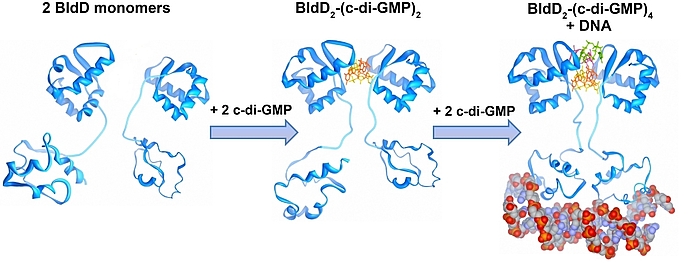
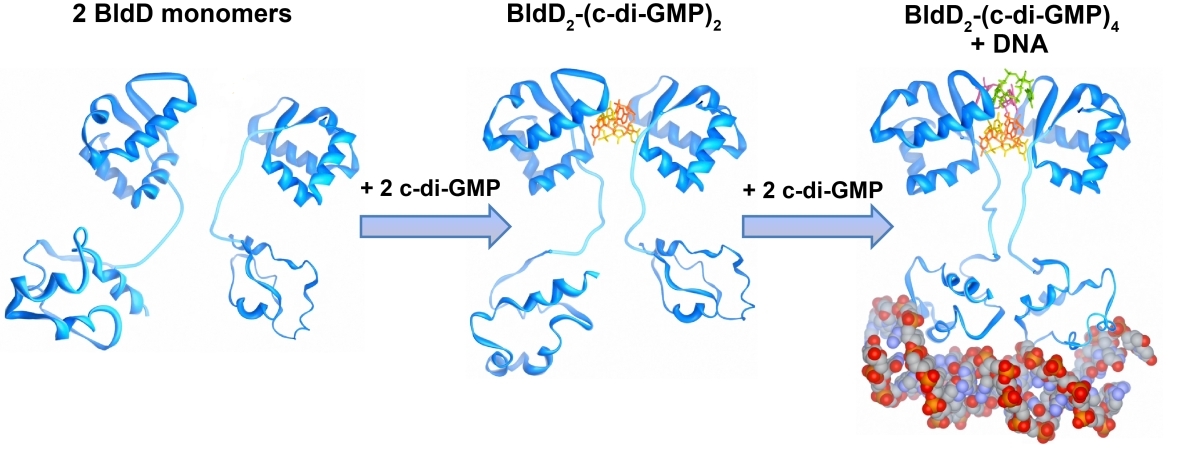
Die Synthese von c-di-GMP aus zwei GTP Molekülen katalysieren Diguanylatzyklasen (DGCs) mit einer GGDEF Domäne, während die Hydrolyse Phosphodiesterasen (PDEs) übernehmen, die durch eine EAL oder HD-GYP Domäne charakterisiert sind. Streptomyces venezuelae, unser Forschungsmodel, besitzt zehn Enzyme, die den intrazellulären Spiegel von c-di-GMP kontrollieren. In unseren derzeitigen Projekten haben wir uns folgende Ziele gesetzt:
- (I) Molekulares und mechanistisches Verständnis der einzelnen c-di-GMP-metabolisierenden Enzyme in der Differenzierung von Streptomyces
- (II) Detaillierte Einsichten in c-di-GMP Verteilungsmustern in zeitlichen und räumlichen Dimensionen in Streptomyces
- (III) Entdeckung von neuartigen c-di-GMP Effektoren und Funktionen in Streptomyces
Förderung: Unsere Forschung zu c-di-GMP in Streptomyces wird vom Emmy-Noether-Programm der Deutschen Forschungsgemeinschaft (DFG) gefördert.
Enzyme, Effektoren und Funktionen von zyklischem di-AMP in Streptomyces
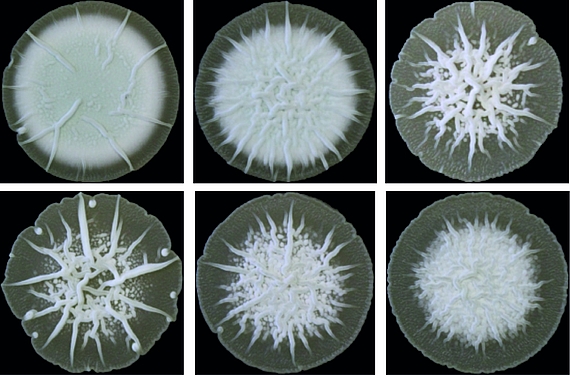
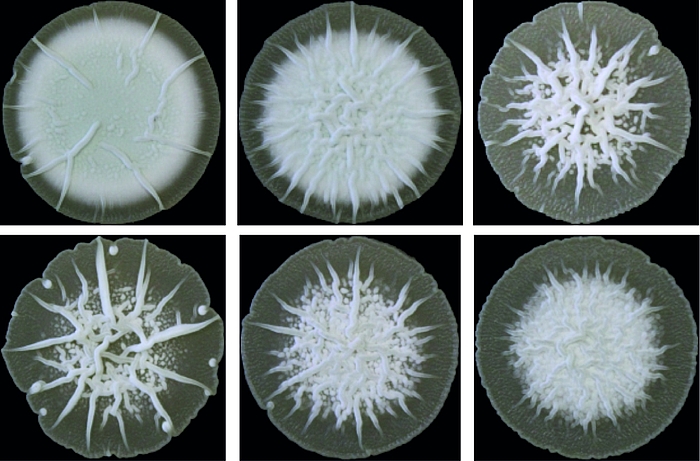
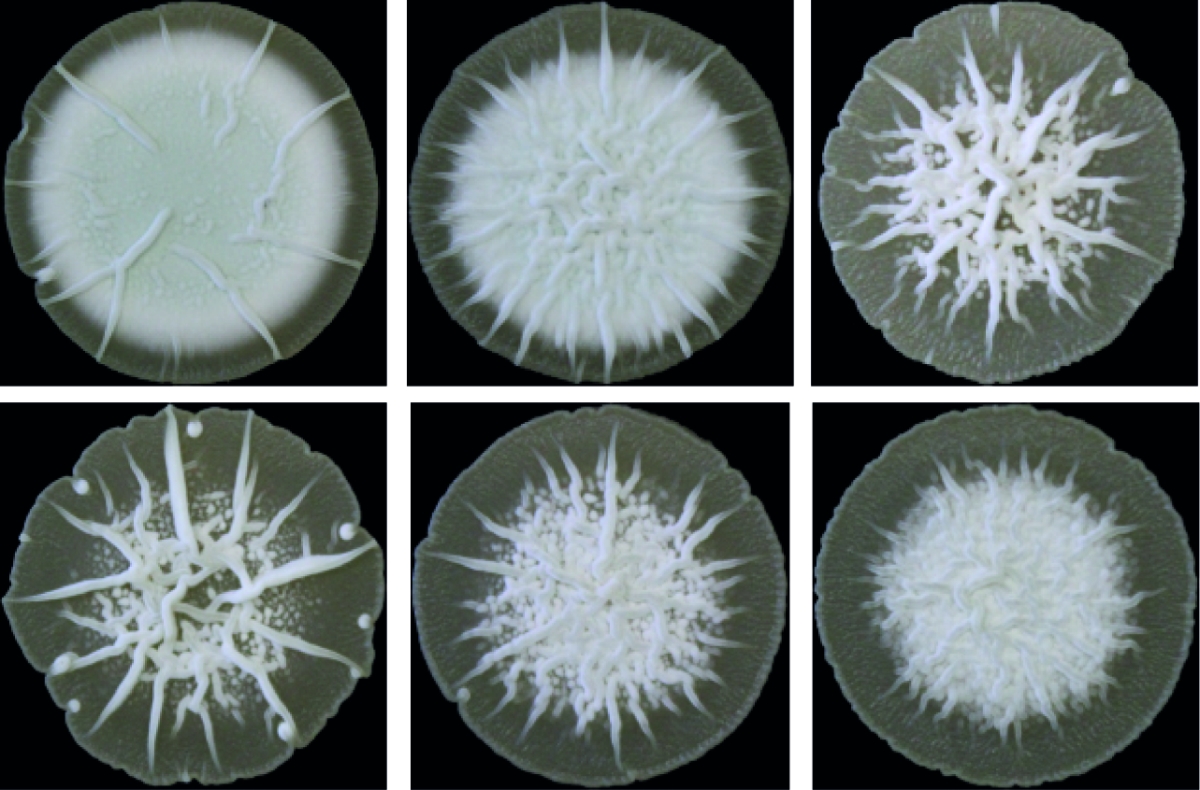
Zyklisches di-AMP ist in Gram-positiven Bakterien und Archeen weit verbreitet und stellt einen wichtigen Regulator der bakteriellen Strategien zur Bewältigung von osmotischen Stress dar. In ihrer Umwelt müssen die Bakterien häufigen Schwankungen der externen Osmolarität standhalten, die zu Zelldehydrierung oder Platzen führen können, wenn adäquate Stressantworten ausbleiben. Als ein wichtiger Bestandteil dieser Mechanismen kontrolliert c-di-AMP den Transport von osmotisch wirksamen Substanzen wie Ionen oder kompatiblen Soluten.
Unser Verständnis zur Funktion von c-di-AMP in Streptomyceten und anderen Aktinobakterien ist sehr oberflächlich. Es war lange unbekannt, wie die Mehrheit der Aktinobakterien c-di-AMP-vermittelte Signale terminieren und welche Effektor Proteine zum Einsatz kommen.
In unseren Studien haben wir das AtaC Protein als eine c-di-AMP-spezifische Phosphodiesterase identifiziert und konnten zeigen, dass das RCK_C-Domänen Protein, CpeA, c-di-AMP bindet. Basierend auf phänotypischen Analysen wissen wir inzwischen, dass c-di-AMP in Streptomyceten die Anpassung an Osmostress und Differenzierungsprozesse reguliert. Die zugrunde liegenden Mechanismen sind jedoch gänzlich unbekannt. Derzeit sind wir auf folgende Fragestellungen fokussiert:
- (I) Welche Komponenten verknüpfen c-di-AMP Signale mit Entwicklungsprozessen in Streptomyces?
- (II) Wie sind Osmostress und Membranintegrität in Streptomyces reguliert?
- (III) Welche Signale lösen c-di-AMP-kontrollierte Vorgänge in Streptomyces aus?
Förderung: Unsere Forschung zu c-di-AMP in Streptomyces wird vom SPP 1879 Schwerpunktprogramm der Deutschen Forschungsgemeinschaft (DFG) gefördert.
SecMess-Funktionen
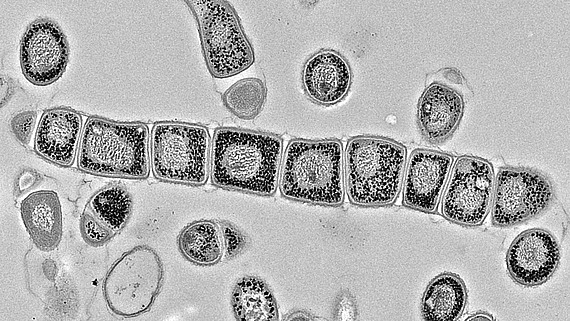


Zur Überwindung der aktuellen globalen Krise der Antibiotikaresistenz, müssen wir neue Wege zur Entdeckung von Antibiotika wagen. Als Beitrag zu dieser Herausforderung werden wir in SecMessFunctions die Signalbiologie von unseren produktivsten Antibiotikaherstellern – den Streptomyceten – untersuchen. Erst seit Kurzem ist bekannt, dass diese Bakterien ein riesiges Reservoir an stillen Genen für die Naturstoffbiosynthese beherbergen, die externe Stimuli zur Aktivierung benötigen. Umweltsignale werden meist über sekundäre Botenstoffe ins Zellinnere übermittelt. Daher betrachten wir diese Signalmoleküle als entscheidend für die Aktivierung der Gene für die Biosynthese von Naturstoffen.
In diesem Projekt befassen wir uns mit dem Potential der sekundären Botenstoffe als Werkzeuge zur Aktivierung von Naturstoff-Biosynthesewege in Aktinobakterien. Darüber hinaus untersuchen wir die Dynamik des bakteriellen Exoskelett-Umbaus als Stressantwort und analysieren neue Mechanismen in „Inside-Out-Transmembrane-Signalling“. Insgesamt wollen wir mit SecMessFunctions folgende Ziele erreichen:
- Etablierung von neuen Erkenntnissen und Richtungen in der Signalforschung
- Identifizierung von neuen Mechanismen in der Zelldifferenzierung und in der Modulation des Zellwandaufbaus in Bakterien
- Entdeckung von Naturstoffen mit neuer biologischer Aktivität
Förderung: Das Projekt SecMessFunctions wird durch den Europäischen Forschungsrat, ERC Starting Grant gefördert.
Bakterielle Multizellularität
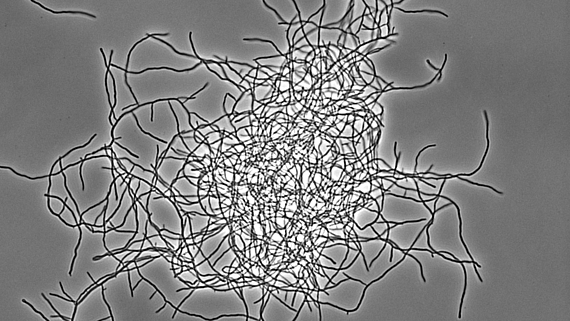

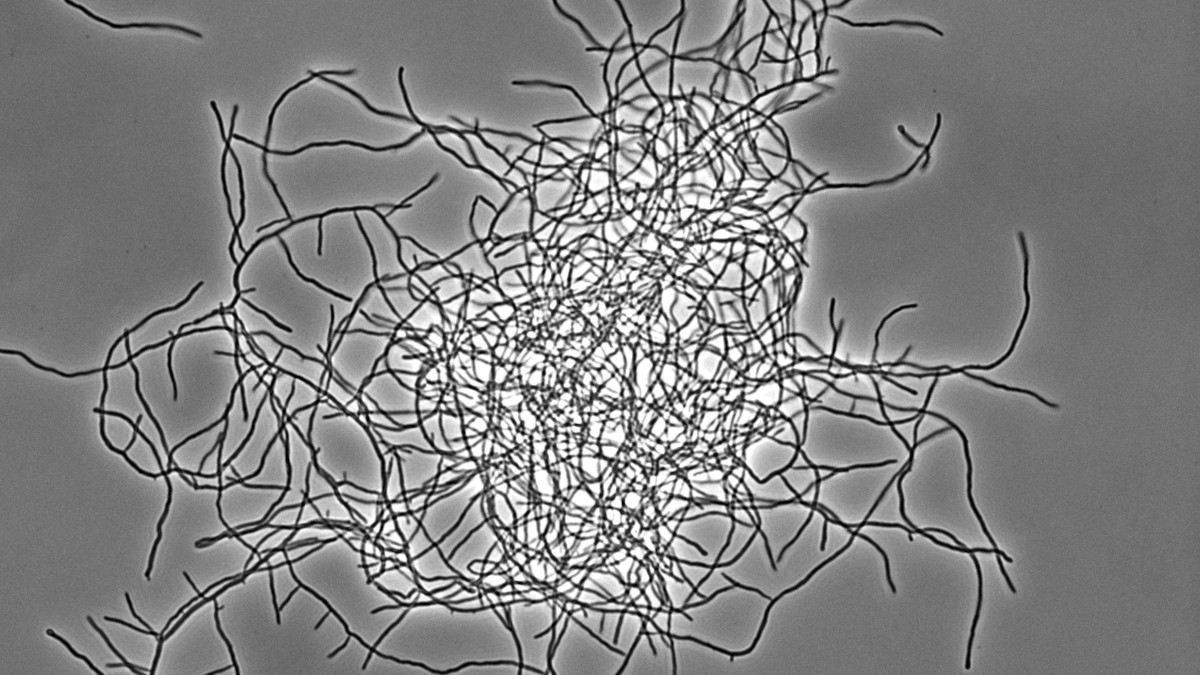
Antibiotika-produzierende Streptomyceten stellen ein einzigartiges, genetisch manipulierbares System zur Untersuchung der bakteriellen Multizellularität dar. Während ihres vegetativen Wachstums bilden sie vielzellige Filamente aus, die durch Querwände in zusammenhängende Kompartimente geteilt sind. Diese filamentöse Form bedingt die Ausbildung von multihyphaler Myzelpellets, die sich zu Kolonien weiterentwickeln. Koordinierte Arbeitsteilung in Kolonien bestimmt welche Subpopulation die Sporulation zur Reproduktion einleitet und welche Zellfraktion Antibiotika zum Schutz der Nährstoffe produziert, die durch Apoptose freigesetzt werden. Welche molekularen Mechanismen diese mehrdimensionale Multizellularität in Streptomyces kontrollieren und welche evolutionären Vorteile sie mit sich bringt, ist jedoch kaum verstanden.
In diesem interdisziplinären Projekt erforschen wir in enger Zusammenarbeit mit dem experimentellen Evolutionsbiologen, Prof. Dr. Daniel Rozen (Leiden Universität, Niederlande) die molekularen Ursachen sowie die Folgen der multizellulären Form in Streptomyces. Wir verfolgen zwei wesentliche Ziele:
- Identifizierung der genetischen Faktoren und der molekularen Mechanismen, die die mehrzellige Existenzform bestimmen
- Quantitative Erfassung der evolutionären Folgen der Mehrzelligkeit auf die Fitness, Arbeitsteilung, Nährstofftransport und Ressourcennutzung in Bakterien
Förderung: Unsere Arbeiten zur Multizellularität von Bakterien wird durch das SPP 2389 Schwerpunktprogramm der Deutschen Forschungsgemeinschaft (DFG) gefördert.
Kontakt


30419 Hannover




