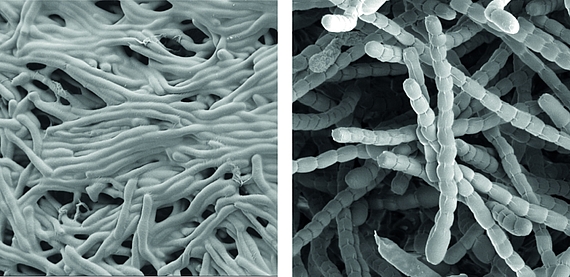
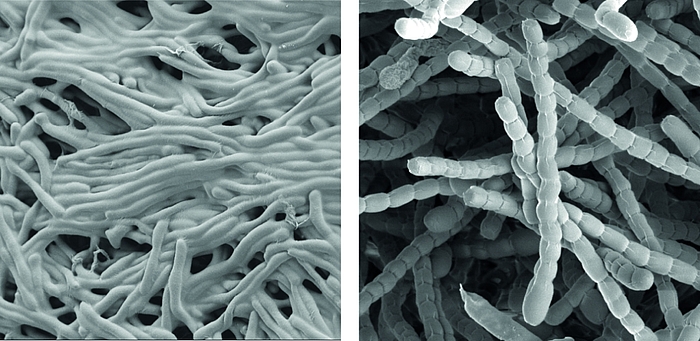
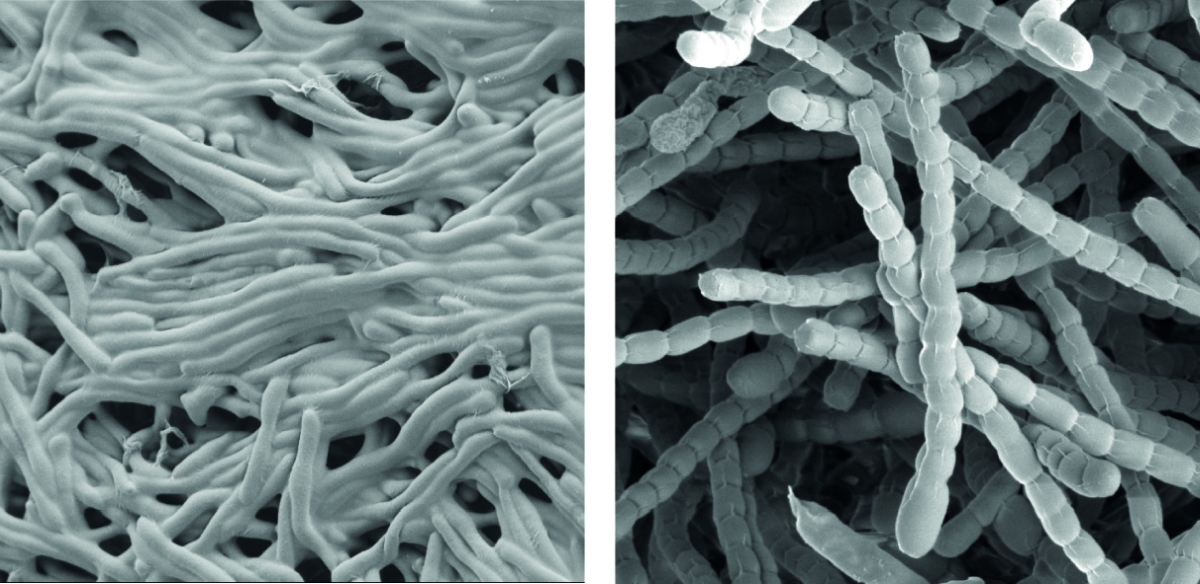
Streptomyces are prominent natural ‘antibiotic factories’ that provide us with the majority of antibiotics used in medicine today.
In addition, they represent an exciting model to study cell differentiation and bacterial multicellularity. During their life cycle, they undergo a complex transition from filamentous, multicellular, vegetative hyphae to unicellular, dormant and highly resistant spores.
Our team
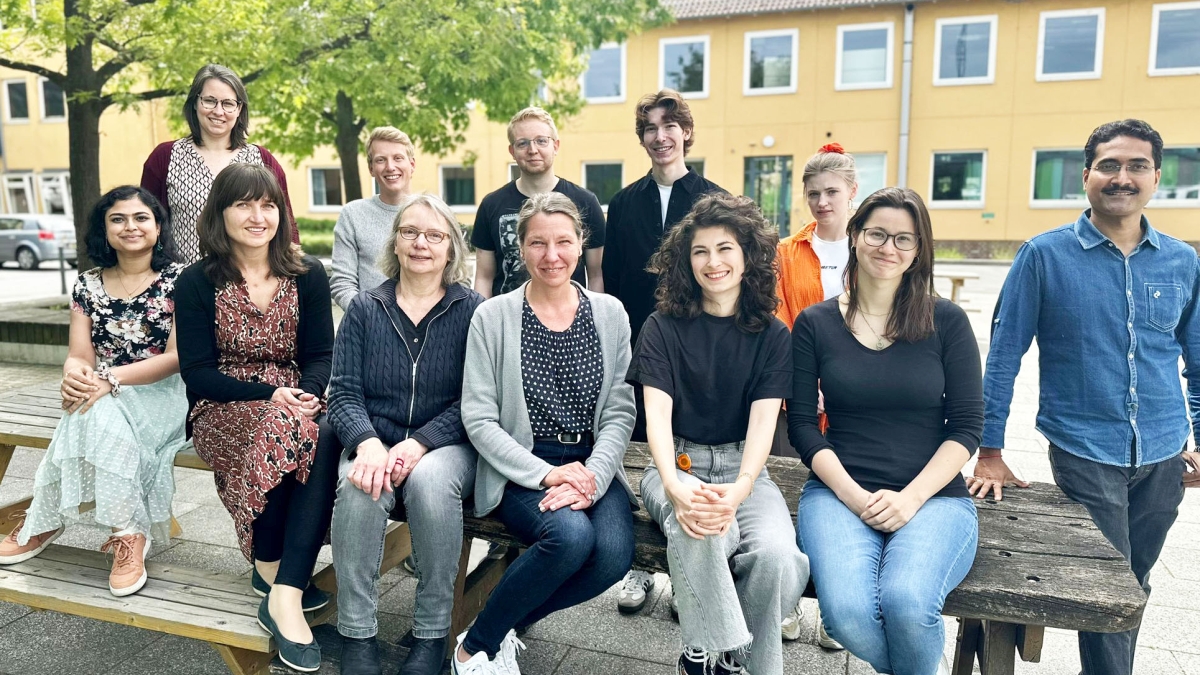



Are you interested in becoming part of our team?
-
Information for postdocs, doctoral candidates or Master's students
We are always happy to welcome new postdoctoral researchers, PhD students or master students in our international research team.
Are you passionate about fundamental biological questions in the area of
- Signal transduction,
- Stress adaptation,
- Cell differentiation,
- Natural products and specialized biopolymers,
- Bacterial multicellularity,
- Regulatory networks and/or
- Streptomyces?
Do you have a background in microbiology, genetics, cell biology or biochemistry? Then please send Natalia an email (tschowri@ifmb.uni-hannover.de) including your CV and a letter of motivation. Funding opportunities often arise on short notice, so initiative applications are more than welcome!
Research projects
Cyclic di-GMP signalling and regulation in bacterial cell differentiation
Cyclic di-GMP is a ubiquitous bacterial second messenger that is renowned for its function in guiding the transition between motility and sessility in most bacteria. In the non-motile streptomycetes, c-di-GMP is a key factor controlling the switch between their filamentous lifestyle and spore formation. Elevated levels of c-di-GMP bring the developmental program to a halt and arrest the cells in the vegetative, filamentous growth phase. Low levels of the cyclic dinucleotide significantly shorten the life cycle by enabling the vegetative hyphae to enter the sporulation programme in a precocious manner. We discovered that the master regulator of Streptomyces development, BldD, is a c-di-GMP-effector that mediates c-di-GMP signals. Binding of c-di-GMP to BldD induces protein dimerization and stimulates BldD-binding to target DNA leading to repression of sporulation genes.
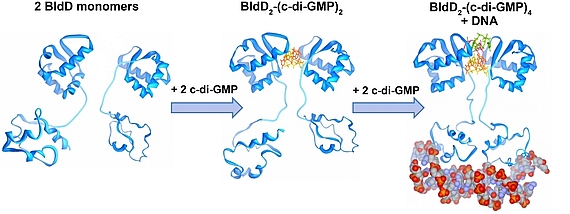
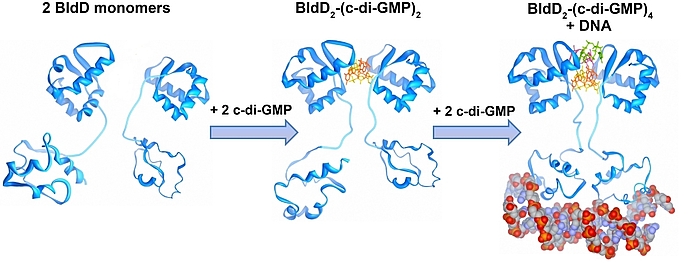
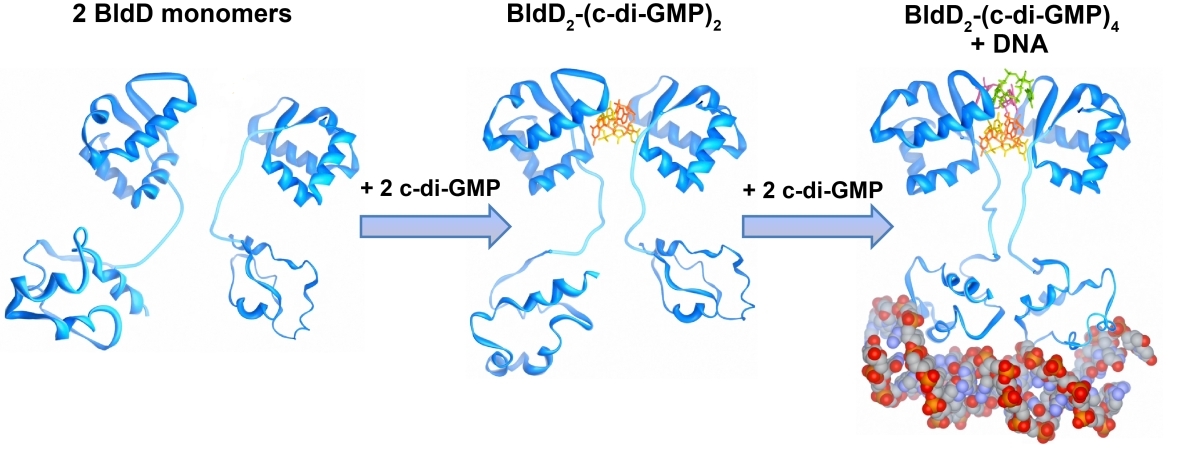
Cellular levels of c-di-GMP are controlled by the competing activities of GGDEF-domain containing diguanylate cyclases (DGCs) that produce the molecule out of GTP, and by phosphodiesterases (PDEs) that carry an EAL or HD-GYP domain to degrade the second messenger. Streptomyces venezuelae, our main lab model, has 10 chromosomally-encoded GGDEF/EAL/HD-GYP genes. In our current projects, we focus on:
- (I) dissecting the physiological functions of individual c-di-GMP enzymes in the differentiation of Streptomyces
- (II) understanding of c-di-GMP patterns in time and space in multicellular Streptomyces
- (III) discovery on novel c-di-GMP effectors and functions in Streptomyces
Funding: Our research on c-di-GMP in Streptomyces is funded by the Emmy Noether Programme from the Deutsche Forschungsgemeinschaft (DFG).
Cyclic di-AMP enzymes, effectors and functions in Streptomyces
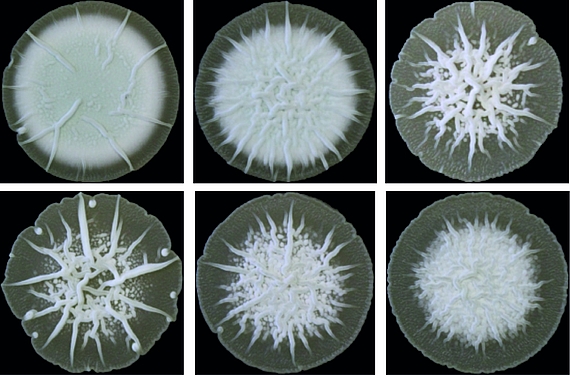
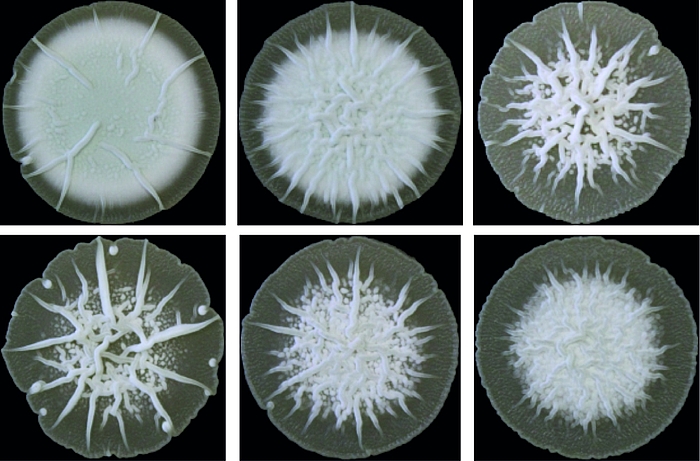
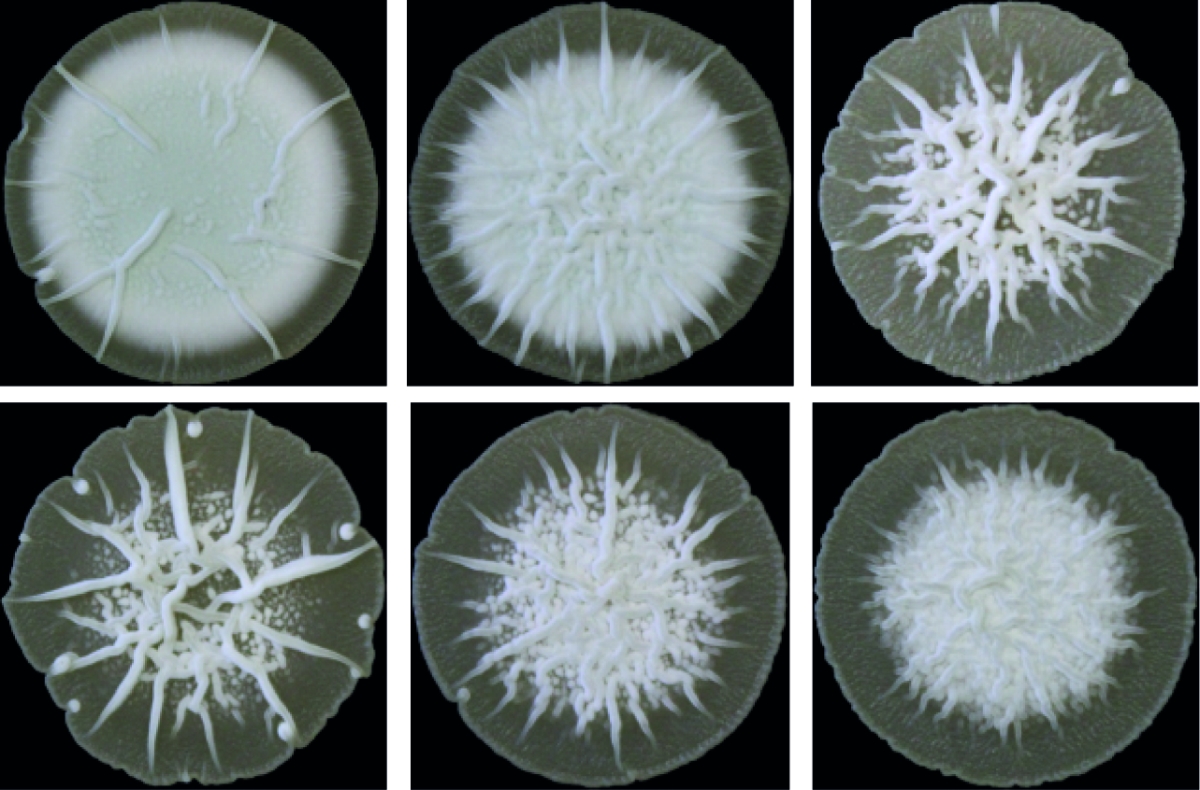
Cyclic di-AMP is mainly produced by Gram-positive bacteria and archaea. Its core function is to control cellular integrity by setting homeostasis of osmolytes that in many bacteria are used for osmoregulation. Changes in external osmolarity trigger water fluxes across the membrane, which can lead to cell dehydration or swelling and, finally, collapse or burst when osmobalance mechanisms fail to respond properly. As a key component of these mechanisms, c-di-AMP directly targets transport systems for osmoactive and osmoprotective substances such as potassium ions and low-molecular-weight compatible solutes in many bacteria.
The function of c-di-AMP in Streptomyces and other Actinobacteria is hardly understood. It was long unknown how the majority of Actinobacteria, stop c-di-AMP signals and which proteins bind the molecule to elicit cellular responses. We recently discovered the AtaC protein as a c-di-AMP-specific phosphodiesterase that is highly conserved in Actinobacteria and the RCK_C domain protein CpeA as a c-di-AMP effector. However, while we know that c-di-AMP in involved in regulation of osmotic stress and development in Streptomyces, the underlying mechanisms are unknown. Currently, we aim at addressing the following questions:
- (I) What are the key components linking c-di-AMP signalling and developmental processes?
- (II) How are osmostress and membrane integrity regulated in Streptomyces?
- (III) Which signals trigger c-di-AMP dependent responses?
Funding: Our research on c-di-AMP in Streptomyces is funded by the SPP1879 Priority Programme from the Deutsche Forschungsgemeinschaft (DFG).
SecMessFunctions
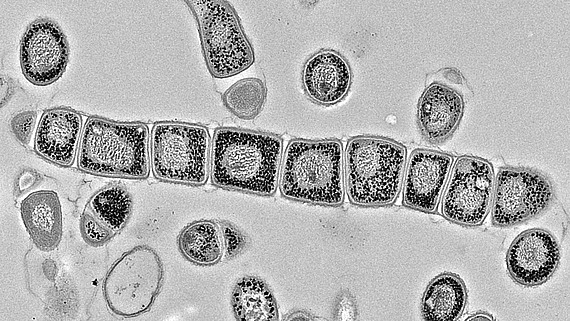


To overcome the current global crisis of antimicrobial resistance, we need to explore new avenues for the discovery of anti-infectives to combat the pathogens. As a contribution to this challenge, in SecMessFunctions we will investigate the signalling biology of our most prolific antibiotic producers – Streptomyces. Only recently we realized, that these bacteria harbour a huge reservoir of silent genes for natural product biosynthesis which require external stimuli for activation. Environmental signals are communicated to the interior of the cell by second messengers. Therefore, we consider these signalling molecules critical for activation of the biosynthesis genes.
In this project, we address the potential of second messengers in activating silent biosynthetic pathways in Actinobacteria. Furthermore, in SecMessFunctions we study the dynamics of bacterial exoskeleton remodelling during stress response and new mechanisms in ‘inside out transmembrane signalling’. Altogether, in SecMessFunction we plan to achieve the following goals:
- Establish new insights and directions in signalling research
- Identify new mechanisms in cell differentiation and cell wall remodelling
- Discover natural products with new biological activity
Funding: The SecMessFunctions project is funded by the European Research Council, ERC Starting Grant.
Bacterial multicellularity
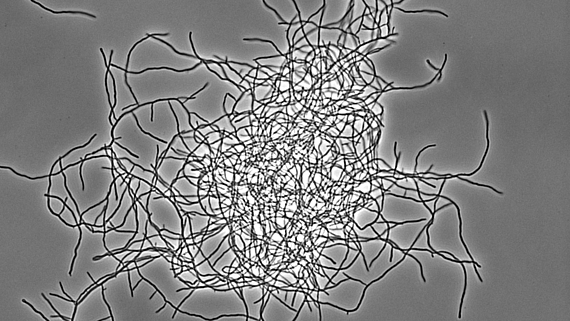

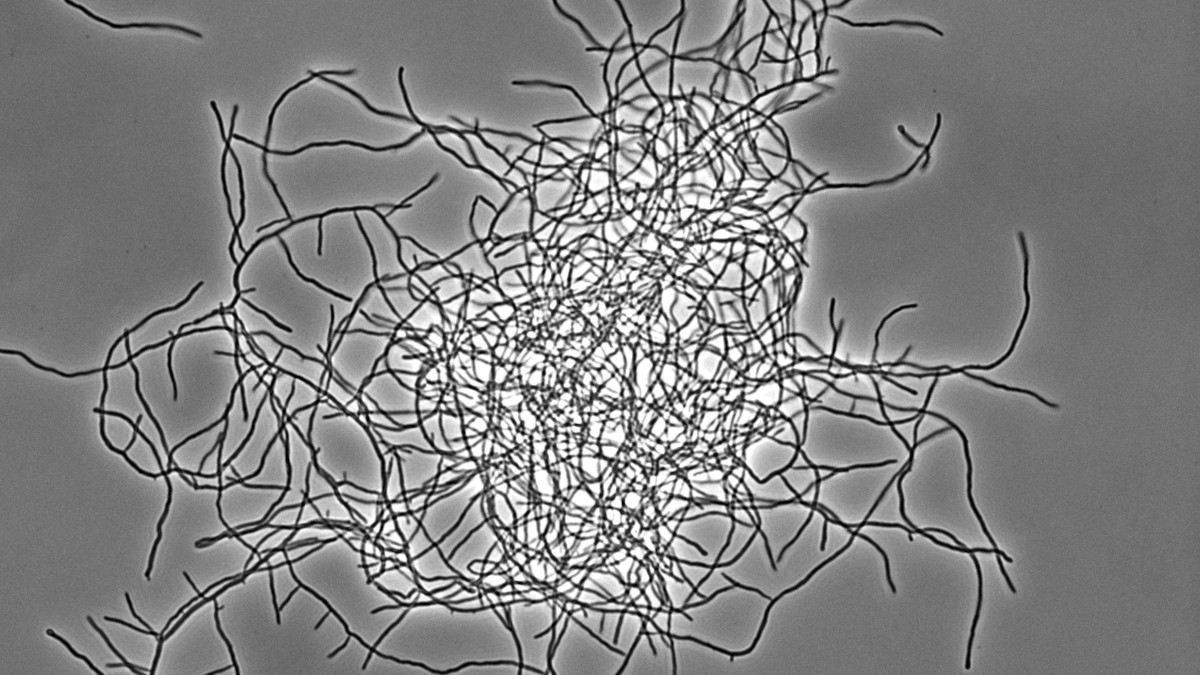
Streptomyces represent an excellent, genetically tractable system to study bacterial multicellularity. During their vegetative growth, they exist as a network of interconnected multicellular filaments containing cross-walls that separate the hypha into compartments but do not lead to cell fission. The filamentous form gives rise to the formation of multi-hyphal mycelial pellets that further develop into structured colonies in which division of labour determines which subpopulation initiates sporulation for reproduction and which fraction of cells produces antibiotics for nutrient protection released by coordinated apoptosis. However, the molecular mechanisms and the evolutionary benefits driving the multi-dimensional multicellularity in the genus Streptomyces are not well understood.
In this interdisciplinary project, in close collaboration with the evolutionary biologist Prof. Dr. Daniel Rozen (Leiden University, Netherlands) we study the molecular factors that determine Streptomyces multicellularity and analyse the fitness benefits of transitions between unicellular and multicellular morphologies. To understand the mechanistic and evolutionary dynamics of multicellular transitions we focus on two major aims:
- Revealing the genetic factors and molecular mechanisms that evolved to control transitions in multicellularity
- Quantify evolutionary consequences of multicellular transitions on fitness, division of labour, nutrient transport and resource use efficiency.
Funding: Our research on Streptomyces multicellularity is funded by the SPP 2389 Priority Programme from the Deutsche Forschungsgemeinschaft (DFG).
Contact


30419 Hannover




